The formulation of the lysis buffer will depend on the subcellular location of the protein of interest, as well as on whether the antibody’s epitope is still present on the protein once it has been denatured. A specific protein of interest is then detected on the membrane using antibodies that recognize an epitope in the target protein, horseradish peroxidase (HRP)-conjugated secondary antibodies, and a chemiluminescent-based detection substrate. The proteins are then transferred to a solid-phase membrane. The sample is then subject to gel electrophoresis allowing the proteins in the mixture to be separated according to their size. In this example, a lysate sample is prepared from cultured cells. Western blotting relies on the electrophoretic separation of proteins from a complex mixture based on their mass, the transfer of these proteins to a solid matrix, and the detection of specific proteins of interest on the matrix using antibodies.įigure 1: The flow-through of a typical western blot experiment. The western blot, also commonly known as immunoblot, has since become an essential and ubiquitous technique in biology and medical labs around the world. The research groups of George Stark at Stanford University and Harry Towbin at Friedrich Miescher Institut in Switzerland published similar immunoblotting techniques at roughly the same time. Despite this, the paper was widely circulated and eventually published in 1981. Interestingly, the manuscript detailing this technique was initially rejected, with reviewers criticizing the method’s name as “flippant and frivolous whimsy”. He subsequently termed this method the “western blot”, in a nod to its predecessors. Neal Burnette, a postdoc at the Fred Hutchinson Cancer Research Center, developed a method for visualizing proteins separated by SDS-PAGE using monoclonal antibodies. This method was quickly followed two years later by the invention of the “northern blot”, which could detect specific RNA molecules using radio-labeled DNA probes. In 1975, Edwin Southern invented the eponymously-named “southern blot”, a technique in which DNA fragments are separated through electrophoresis based on their size and then transferred to a nitrocellulose membrane for detection. In fact, essential western blot methodology, including sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and monoclonal antibodies with antigen specificity, were only established in 19, respectively. 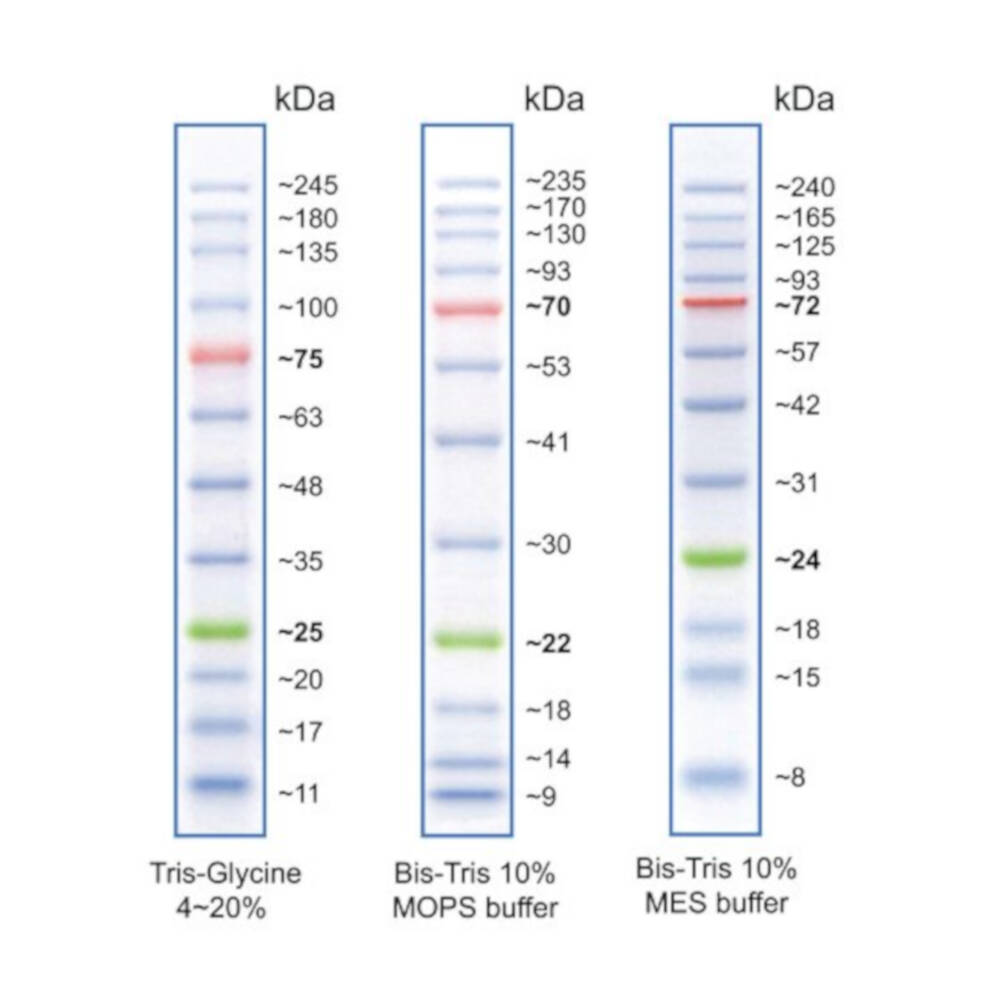
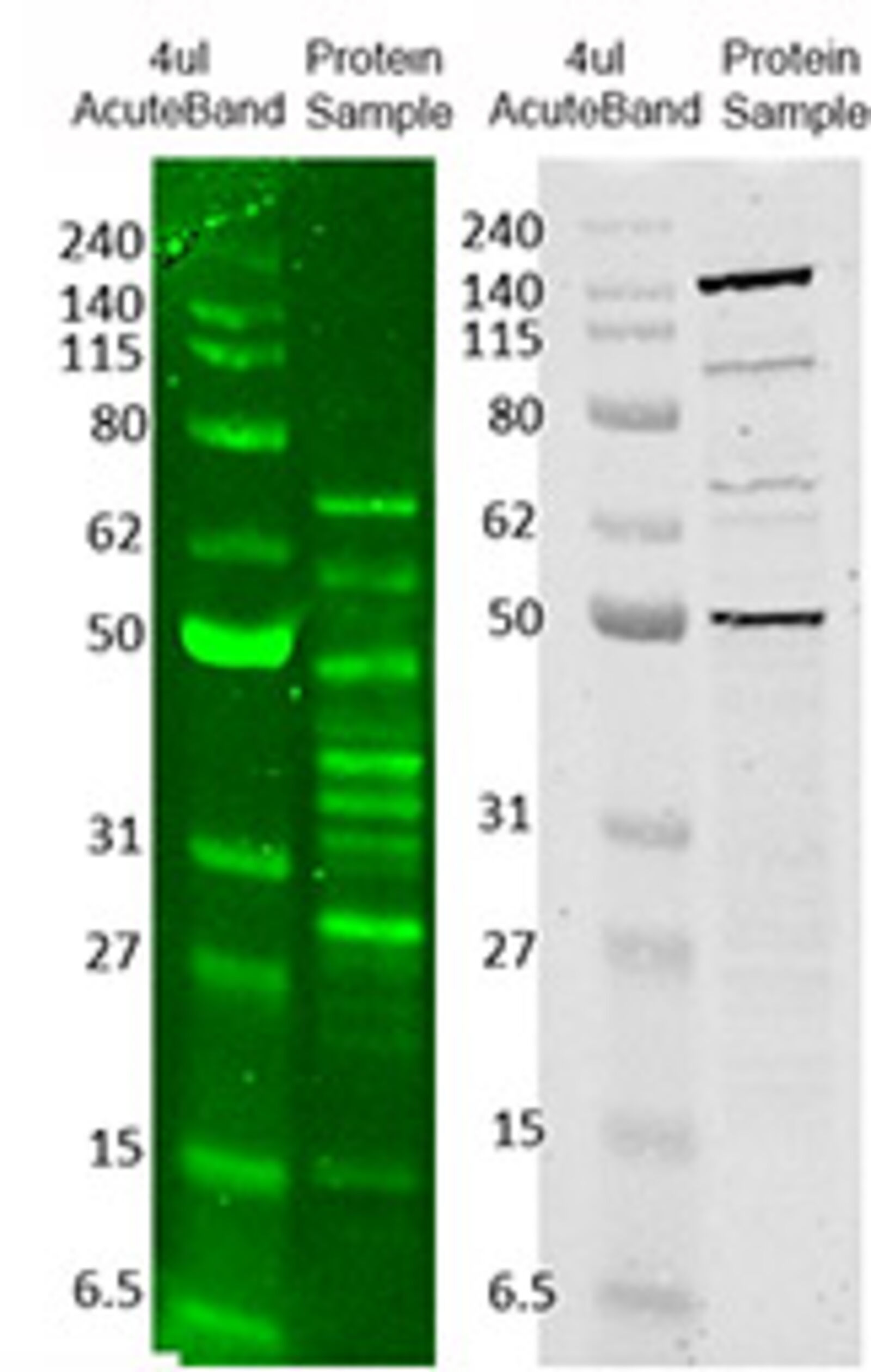
Protein molecular weight markers are sometimes used as a control when verifying protein migration and observing transfer efficiency.It may be surprising to learn that the history of the western blot only stretches back to the late 1970s. They are most commonly used in SDS-PAGE and western blotting. 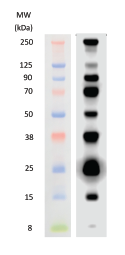
Protein molecular weight markers consist of proteins whose weight is measured in kDa. Because RNA migrates faster than DNA through electrophoresis gels, the appropriate nucleic acid ladder (RNA vs DNA) should be used. To determine fragment sizes, RNA is typically run in a denaturing electrophoresis gel. Similarly to DNA ladders, RNA ladders contain RNA fragments of varying sizes. They are great when more specific fragment length verification is desired than can be inferred from a typical ladder with larger, or less consistent increments. Step Ladders are not intended for concentration estimations. For example, step ladder DNA fragments could be in increments of 100bp from 100bp all the way up to 4000bp. Often, DNA molecular weight markers and ladders come in two formats: pre-mixed with loading dye, or with the loading dye separate.ĭNA Step Ladders contain DNA bands of defined sizes and equal intensity with fragment lengths at defined increments. These ladders are sometimes made from restriction enzyme digestion of plasmid DNA with a known sequence. When the markers are loaded in adjacent lanes to samples, sample fragment or protein size and concentration estimations are easily determined.ĭNA ladders contain DNA fragments of varying sizes. These standards contain pre-determined fragment (or protein) sizes and concentrations. Molecular weight markers, or ladders, are a set of standards that are used for determining the approximate size of a protein or a nucleic acid fragment run on an electrophoresis gel.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
